Revolutionary approach for drug-resistant TB has important implications for other diseases
Published 31 January, 2022
Tuberculosis (TB) is a potentially fatal bacterial disease that typically affects the lungs. According to the World Health Organization (WHO) and Centers for Disease Control and Prevention (CDC), there are 9-10 million new TB cases each year, resulting in 1-2 million deaths. In fact, TB is one of the top 10 causes of death globally.
Researchers have previously discovered four first-line drugs that treat mycobacterium tuberculosis (Mtb), the bacteria causing TB. These include ethambutol (EMB), which typically blocks Mtb membrane synthesis. However, around 4% of TB cases have genetic mutations that make them drug resistant.
Previous studies have identified that some of these cases are caused by EmbR mutations binding with Mtb PknH, a transmembrane kinase. However, researchers have been unable to identify the exact binding site and potential mechanisms of the Mtb mutation.
In a pioneering study, published in the KeAi journal Synthetic and Systems Biotechnology, scientists used a revolutionary approach in computer-aided drug discovery to explore binding sites between resistance-causing mutations of EMB, and a transmembrane kinase. It is the first study to propose this protein-protein interaction site and its binding partner based on its disordered protein nature.
According to senior author Prof. Gil Alterovitz, of the Harvard Medical School/ Brigham and Women's Hospital in the US, “this kind of computer-aided drug-discovery has the potential to improve the efficiency of discovering new binding sites and binding partners, and aid in the development of many other critical treatments for world-threatening diseases.”
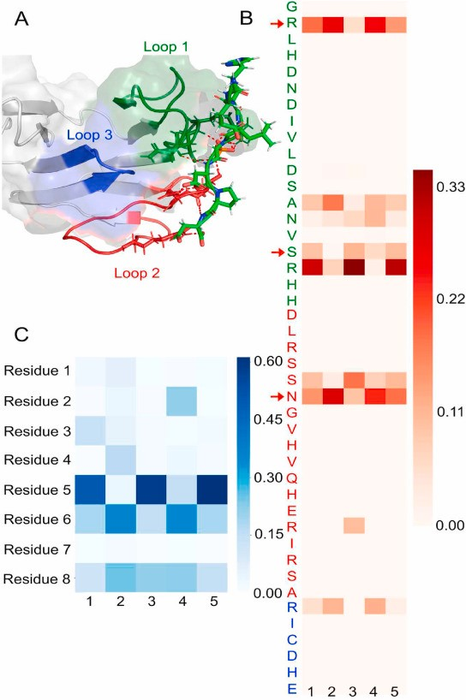
By elaborating findings from a 2006 protein study by Alderwick LJ, et al., the international team determined and investigated the structure of a specific binding site which showed finger-like structures holding the phosphorylated residues displayed looser binding. Hydrogen bond analysis and determination of the C-terminal backbone torsional angles supported the peptides’ binding towards EmbR and involvement in phosphorylation. This phosphorylation can cause structural changes, therefore impacting its overall function.
Prof. Alterovitz adds: “The study’s novel approach empowers new ways of thinking on drug resistance and novel mechanisms for dealing with it. The findings have the potential to revolutionise not only the study of drug-resistant Mtb treatments, but also the field of computer-aided drug discovery at-large.”
###
Contact the corresponding author: Gil Alterovitz, gil_alterovitz@hms.harvard.edu